Water. Three years ago, on Rare Disease Day 2023, I published a blog post comparing phenotypes to water, existing in three phases: solid, liquid, and vapor. At the time, I was trying to make a simple point. Rare diseases are not defined by their first description, nor only by a checklist of features. They are moving targets. Here is what has changed since then.
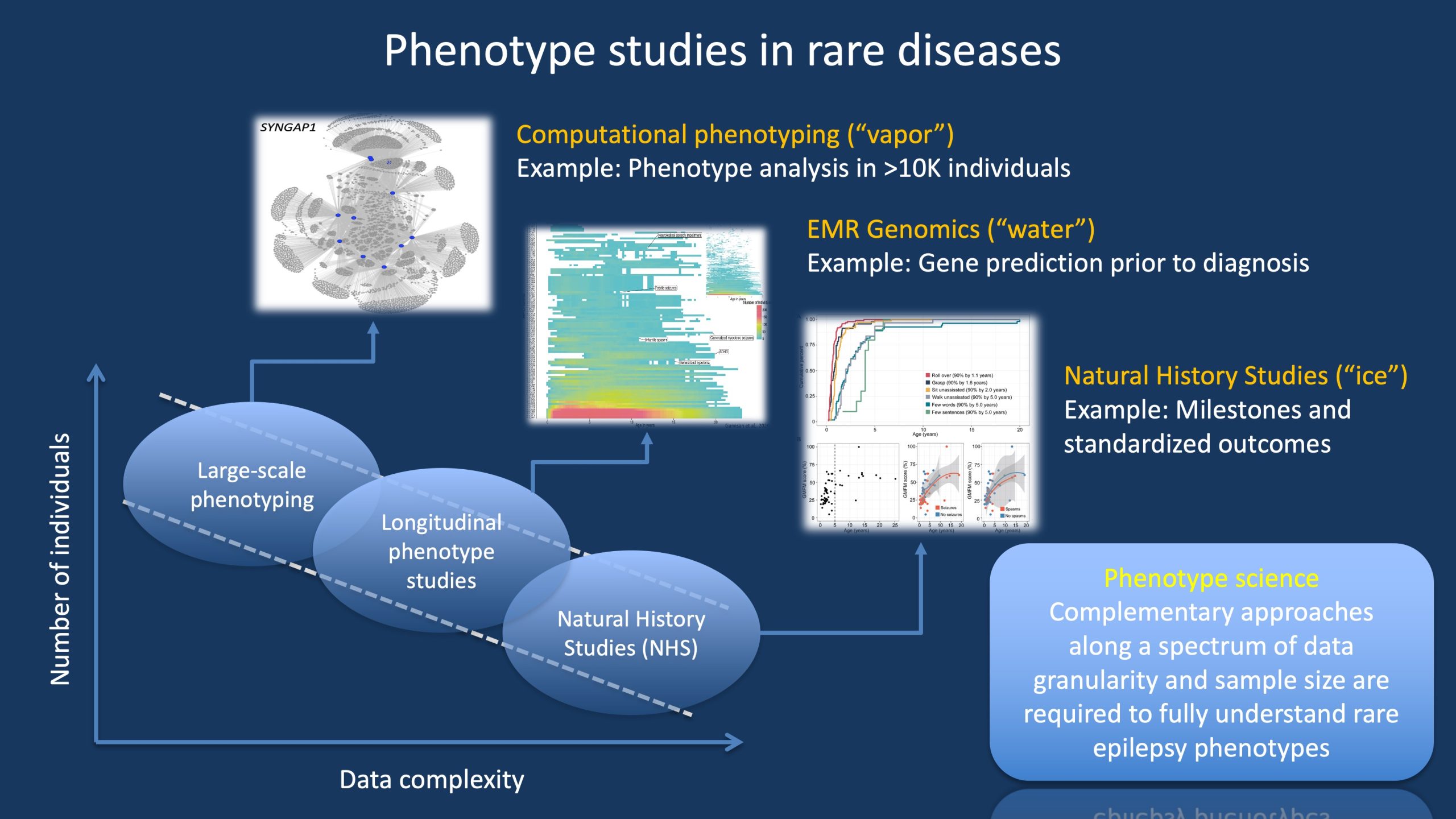
Figure 1. There are different ways to gain understanding of rare disease phenotypes, ranging from approaches with large patient numbers and core phenotypes to smaller studies with highly granular phenotypes. In our blog post, we use the analogy of the different phases of water (ice, water, vapor) to represent the three main approaches to phenotype studies. Importantly, these approaches are not mutually exclusive but complement each other. For example, Natural History Studies and clinical trial readiness studies would be limited if longitudinal phenotype studies were not available to provide a broad overview of the disease trajectory. However, in turn, these longitudinal studies depend on large-scale phenotyping studies to provide an overview of broader disease patterns and subgroups.
Phases of phenotypes. I must admit that I am leaning into an analogy that is less than perfect, but by comparing phenotypes in rare diseases to the three phases of water, I am trying to emphasize that there are different ways we can approach clinical features. These include approaches that are established (ice), more fluid and emergent (water), and innovative approaches that push the boundaries (vapor). They all have their place and contribute different aspects to how we understand phenotypes.
Ice – Natural History Studies. The established, solid-phase approaches to phenotypes are natural history studies (NHS) that can almost become clinical trials in themselves by using FDA concepts and regulatory language. I can only attest to our growing NHS data on STXBP1 (STARR) and SYNGAP1 (ProMMiS), but these datasets are expanding, providing clearer developmental trajectories and better-defined outcome measures. A few weeks ago, I used the terms Minimal Detectable Change (MDC) and Minimal Clinically Important Difference (MCID) in a grant application, and it felt like second nature to handle these concepts. This was unthinkable only three years ago. In addition, biomarkers have made a comeback in genetic epilepsies, most notably quantitative EEG biomarkers. These biomarkers are no longer exploratory add-ons but are part of the expected architecture of interventional studies. At the risk of overextending this comparison, the ice has thickened since 2023.
Water – Real-world data. In 2023, I described EMR genomics as the flowing river beneath the ice. What has changed since then is the widespread adoption of real-world data. We have contributed additional analyses on SYNGAP1, SCN8A, and TBC1D24. Multi-institutional datasets now allow reconstruction of seizure trajectories, medication responses, and comorbidity patterns across thousands of patient-years. Comparative effectiveness signals, such as medication response in specific genetic epilepsies, are increasingly reproducible. The misconception that rare diseases lack data has become harder to defend. The river of fluid phenotypic data has become much wider since 2023.
Vapor – Computational Phenotyping. But there is even more to phenotypes that we need to understand, particularly within the larger framework of how clinical features in genetic epilepsies and neurodevelopmental disorders connect to each other. For most conditions, this information cannot be gathered from natural history studies or real-world data alone. For example, in 2025, we reconstructed the phenotypic spectrum of BSN-related disorders almost entirely from de-identified biobank data and minimal phenotypic information across resources. For this type of analysis, we need a deep understanding of how to connect sparse data points, how to handle absence of information, and how to access data that initially seems untraceable. We need to develop algorithms and understand what they are doing. However, this approach allows us to address the large proportion of rare diseases that are currently unmapped. We are now more confident that we can reconstruct recognizable syndromes and delineate variant-specific subgroups. Pattern recognition at scale has become routine and guides gene discovery and disease delineation.
Phase shift to data integration. Today, the biggest change is not within any single phase, but in how this information fits together. I predict that over the next three years, we will see an increasing focus on data integration. Trial readiness studies can draw on EMR-derived trajectories, and large-scale computational phenotyping can inform which subgroups warrant deeper natural history studies. Work in these three fields is often still unconnected, as it cuts across traditional genomic research, EMR analysis, and clinical studies. But we need all three phases of phenotypes to move forward.
What you need to know. Three years ago, I argued that phenotypes behave like water. We can understand clinical features on at least three different levels that differ in depth and scope (Figure 1), just like ice, water, and vapor. If we want to move toward targeted therapies, we need to understand phenotypes on all three levels.
