qEEG. The electroencephalogram is one of the oldest tools in neurology. We use it every day to diagnose and monitor brain function, yet, even in the era of genomic medicine, most of our EEG interpretation still relies on visual inspection, a human reading of squiggled traces. In a recent publication in Neurology, we asked whether the information embedded in these signals could be measured more objectively in children with STXBP1, SCN1A, and SYNGAP1-related disorders. Here is the story on how we identified hidden signals in the EEG tracings of individuals with genetic epilepsies.
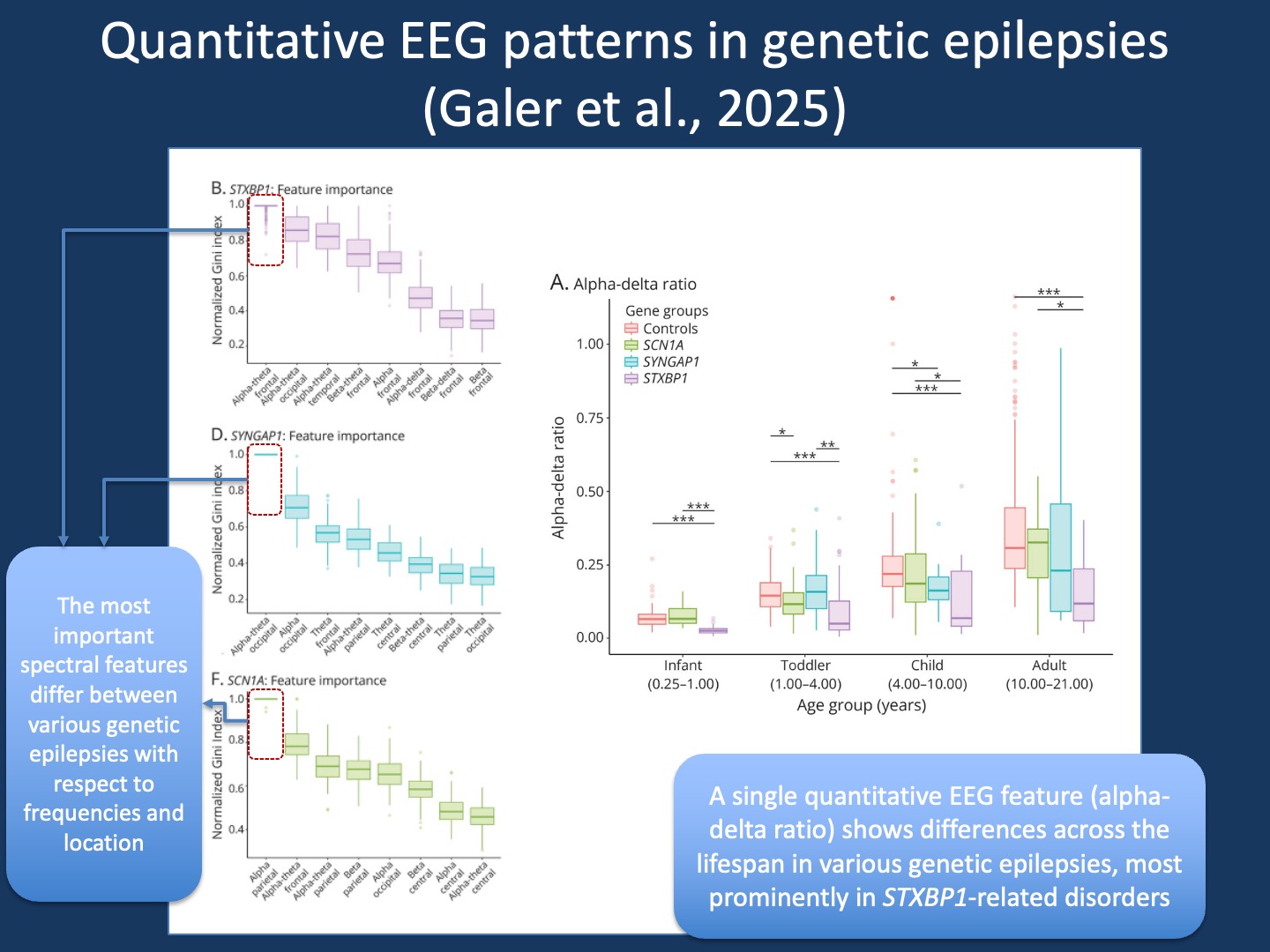
Figure 1. Quantitative EEG features in genetic epilepsies. Using machine learning, we were able to extract specific qEEG features that were more most important in separating controls from individuals with STXBP1-, SYNGAP1-, and SCN1A-related epilepsies (feature importance, left). The global alpha-delta ratio is a previously studied qEEG feature. When assessing this across age groups and various genetic epilepsies, we saw that this measure is most prominent in STXBP1-related disorders across all age groups, while it was only significantly lower than controls in other conditions (e.g. SCN1A) at specific ages.
Turning EEG into data. The project grew from a simple fact: every clinical EEG is already digital and can, in principle, be analyzed again in the future. Led by Peter Galer as part of his PhD thesis, we built an automated pipeline that could clean, segment, and quantify thousands of EEGs from the electronic medical records. From these data, we extracted spectral features—the power allocated to different frequency bands—and asked whether these patterns differ between the major genetic epilepsies and typically developing controls.
EEG as real-world data. All EEGs in this study were recorded during routine clinical care: not in a research laboratory, but in everyday diagnostic settings. This distinction matters. Most importantly this information is readily available and large. We were able to retrieve 154 EEGs for individuals with SCN1A-related disorders, 95 for STXBP1, 46 for SYNGAP1, and a total of 847 control EEGs from 806 individuals without epilepsy. The data reflect the variability, noise, and imperfection of real-world practice, yet they still hold measurable signals that can be extracted at scale. This demonstrates the potential of electronic health record–based research to uncover new biomarkers directly from the clinical workflows that generate them.
From promise to practicality. Quantitative EEG biomarkers hold enormous promise. They can provide objective, reproducible measures of brain function that may capture aspects of disease progression invisible to standard visual EEG inspection. However, obtaining such biomarkers prospectively in rare disease research is both time- and cost-intensive. For this reason, repurposing real-world clinical EEGs offers a pragmatic first step. These retrospective datasets allow us to evaluate how informative specific quantitative measures might be for a given condition before committing resources to implementing this in prospective natural history studies or clinical trial. In this way, clinical data become a proving ground for which biomarkers are most likely to translate into meaningful endpoints.
Distinct electrophysiologic signatures. What emerged from our analysis in our publication by Galer et al were subtle but consistent signatures. For example, individuals with STXBP1-related disorders showed a reduced frontal alpha–theta ratio. Spectral models could distinguish EEGs from STXBP1, SCN1A, and SYNGAP1 with high accuracy, each with a characteristic spatial pattern: frontal features in STXBP1, parietal in SCN1A, and occipital in SYNGAP1. These observations echo what many of us sense clinically, namely that each of these genes affects neural networks in distinct ways. This assumption is now supported by quantitative evidence.
What EEGs can and cannot tell us. We also asked whether these measures might reflect clinical outcomes. In our analysis, taking into account the noisy data from real-world medical records, the EEG spectral features were not predictive of seizure frequency. However, EEG features correlated with gross motor function measured by the GMFM-66, performing better than models based on age alone. This suggests that some aspects of brain organization captured by our quantitative EEG measures may parallel development attainments during childhood. This observation motivated us to assess more closely how qEEG features and developmental outcomes correlate in these conditions, a comparison that requires the high-quality data from prospective natural history studies such as STARR and ProMMiS.
Looking ahead. The larger message of this work is that ordinary clinical data can be repurposed for discovery. We have previously analyzed real-world clinical data collected in the EMR for STXBP1, SYNGAP1, and SCN8A. With our study by Galer and collaborators, we are now able to demonstrate that the utility of real-world data also extends to electrophysiological data. Each EEG recorded at the bedside contains quantitative information that, when analyzed systematically, may add to our understanding about disease trajectories, treatment response, and outcomes.
What you need to know. In our publication by Galer and collaborators in Neurology, we were able to demonstrate that quantitative EEG analysis can uncover distinct electrophysiologic signatures in STXBP1, SCN1A, and SYNGAP1-related epilepsies. While quantitative EEG biomarkers are promising, collecting them prospectively remains resource-intensive. Repurposing clinical EEGs offers a feasible, data-driven starting point to understand which measures truly matter for specific genetic epilepsies and how they might one day serve as reliable endpoints in clinical research.
